From MdsWiki
| Revision as of 13:38, 15 July 2015 (edit) Manduchi (Talk | contribs) (→<center><font face="Arial"><big>More on MDSplus expressions</big></font></center>) ← Previous diff |
Current revision (14:28, 24 May 2023) (edit) Manduchi (Talk | contribs) (→<center><font face="Arial"><big>Understanding MDSplus signals</big></font></center>) |
||
| Line 3: | Line 3: | ||
| | | | | ||
| __NOTOC__ | __NOTOC__ | ||
| + | |||
| + | === General Notation: The Data Hierarchy - Trees, Nodes, and Models === | ||
| + | |||
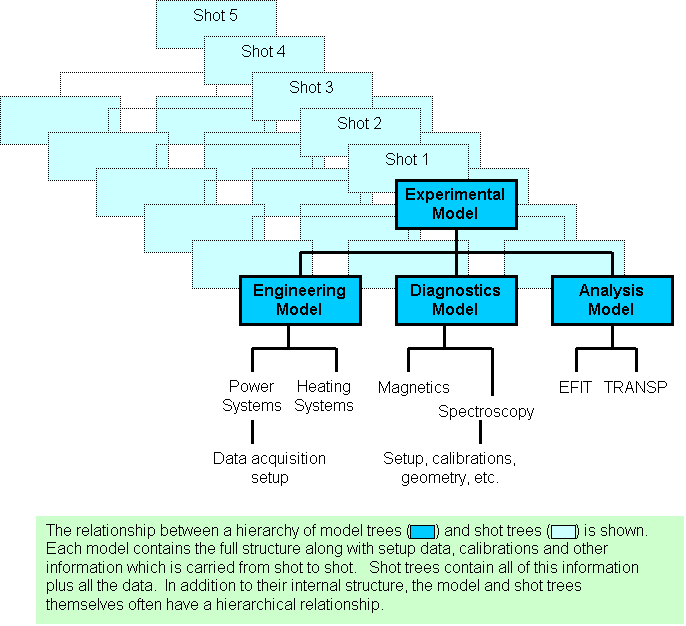
| + | The basic data structure of MDSplus is a self-descriptive hierarchy called a TREE. The hierarchy consists of large numbers of named NODES which make up the branches (structure) and leaves (data) of each tree. MDSplus SHOTS are trees created from a special type of tree called a MODEL, a template which contains all of the structure and setup data for an experiment or code. SHOTS are copies of the model augmented by the stored data and correspond to particular runs of an experiment or code. For a typical experiment, data from various sources are grouped in some logical manner and divided into a number of trees which each form the top level of their respective hierarchies. These trees themselves can be organized into a hierarchy with a root tree and SUBTREES as in the following figure. | ||
| + | |||
| + | [[Image:Tree-with-caption.gif]]<br /><br /> | ||
| + | In the rest of this section we shall discover how MDSplus data structures can be created and accessed. Let's start! | ||
| + | |||
| ===How to create and populate a MDSplus tree=== | ===How to create and populate a MDSplus tree=== | ||
| Line 14: | Line 22: | ||
| ---- | ---- | ||
| + | |||
| === A Quick Check for some important environment variables === | === A Quick Check for some important environment variables === | ||
| Before starting working with MDSplus, it is important to make sure that a few environment variables are properly set. When installed via an install shield or RPM, the variables should be ok, but in the case MDSplus is build from the source distribution, the following variables must be defined for Linux: | Before starting working with MDSplus, it is important to make sure that a few environment variables are properly set. When installed via an install shield or RPM, the variables should be ok, but in the case MDSplus is build from the source distribution, the following variables must be defined for Linux: | ||
| Line 153: | Line 162: | ||
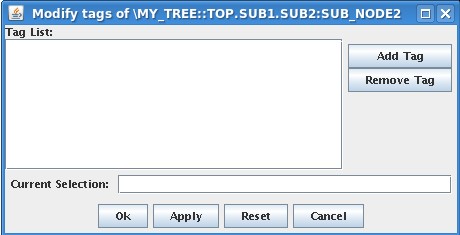
| <center><font face="Arial">[[Image:ModifyTags.jpg]]</font></center> | <center><font face="Arial">[[Image:ModifyTags.jpg]]</font></center> | ||
| - | <font face="Arial">in which you can add/remove the tag name(s) which will be associated with that node. Add MY_SPECIAL_NODE in the list (writung the name in ''Current Selection'' Field and then pressing Add Tag button) and press Ok. From now, the tag is associated with that node, and is shown in the data dialog when showing or modifying data. </font> | + | <font face="Arial">in which you can add/remove the tag name(s) which will be associated with that node. Add MY_SPECIAL_NODE in the list (writing the name in ''Current Selection'' Field and then pressing Add Tag button) and press Ok. From now, the tag is associated with that node, and is shown in the data dialog when showing or modifying data. </font> |
| <font face="Arial">(Note that in this example, the first time you select the '''modify data''' popup item for this node, the node will be shown as undefined, as no data has been added to it in TCL. You can insert new data into it by changing the '''undefined''' option into '''expression''' and then typing an expression)</font> | <font face="Arial">(Note that in this example, the first time you select the '''modify data''' popup item for this node, the node will be shown as undefined, as no data has been added to it in TCL. You can insert new data into it by changing the '''undefined''' option into '''expression''' and then typing an expression)</font> | ||
| Line 235: | Line 244: | ||
| <font face="Arial">Let's add to my_tree a node called SIGNAL1 containing a sine wave made of 1000 sampled points taken over a period of 1 second.</font> | <font face="Arial">Let's add to my_tree a node called SIGNAL1 containing a sine wave made of 1000 sampled points taken over a period of 1 second.</font> | ||
| - | TCL>edit my_tree<br />TCL>add node SIGNAL1/usage=signal<br />TCL>put SIGNAL1 "build_signal(sin(6.28 * [0..999]/1000.),, [0..999]/1000.)"<br />TCL>write<br />TCL>close | + | TCL>edit my_tree<br />TCL>add node SIGNAL1/usage=signal<br />TCL>put SIGNAL1 "build_signal(sin(6.28 * [0 .. 999]/1000.),, [0 .. 999]/1000.)"<br />TCL>write<br />TCL>close |
| <font face="Arial">The TDI expression syntax for defining signals is the following (full reference to TDI expressions is [[Documentation:Reference:TDI|here]] ):</font> | <font face="Arial">The TDI expression syntax for defining signals is the following (full reference to TDI expressions is [[Documentation:Reference:TDI|here]] ):</font> | ||
| Line 243: | Line 252: | ||
| <font face="Arial">where:</font> | <font face="Arial">where:</font> | ||
| - | <font face="Arial"> - DATA is an expression defining the Y values. (sin(6.28 * [0..999]/1000.) in our example)<br /> - RAW is an expression indicating raw data. Often an acquired signal is made of raw data converted then by taking into account parameters such as gain and offset. For the moment we omit its definition (the two commas in the TCL example are not a typing error).<br /> - DIMENSION is an expression returning the array of X axis, usually (but not always) time values. (In our example the expression is [0..999]/1000.)</font> | + | <font face="Arial"> - DATA is an expression defining the Y values. (sin(6.28 * [0 .. 999]/1000.) in our example)<br /> - RAW is an expression indicating raw data. Often an acquired signal is made of raw data converted then by taking into account parameters such as gain and offset. For the moment we omit its definition (the two commas in the TCL example are not a typing error).<br /> - DIMENSION is an expression returning the array of X axis, usually (but not always) time values. (In our example the expression is [0 .. 999]/1000.)</font> |
| <br /> | <br /> | ||
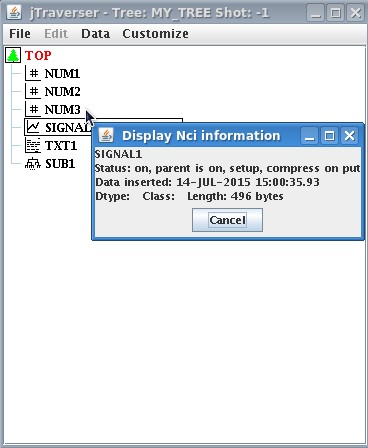
| Now let's check the dimension of the created data item. One may expect that the signal just created will require the bytes necessary to store 1000 float values for the Y value plus 1000 float values for the times. Assuming that 4 bytes are reserved for a floating point number, this yields 8KB. We can inspect the dimension of SIGNAL_1 data item using the popup menu option '''Display Nci'''. The window below is shown | Now let's check the dimension of the created data item. One may expect that the signal just created will require the bytes necessary to store 1000 float values for the Y value plus 1000 float values for the times. Assuming that 4 bytes are reserved for a floating point number, this yields 8KB. We can inspect the dimension of SIGNAL_1 data item using the popup menu option '''Display Nci'''. The window below is shown | ||
| [[Image:SignalSize.jpg]] | [[Image:SignalSize.jpg]] | ||
| showing that the dimension of the stored item is 496 bytes!!! How can it be?? The reason is that what has been stored in the tree is an expression specifying a sinusoidal signal, '''NOT''' the points of the sinusoidal signal itself. However, whenever accessed, such expression will be evaluated on the fly by MDSplus, thus producing the 1000 float values for the Y and X axis, respectively. <br /><br /> | showing that the dimension of the stored item is 496 bytes!!! How can it be?? The reason is that what has been stored in the tree is an expression specifying a sinusoidal signal, '''NOT''' the points of the sinusoidal signal itself. However, whenever accessed, such expression will be evaluated on the fly by MDSplus, thus producing the 1000 float values for the Y and X axis, respectively. <br /><br /> | ||
| - | This may seems a bit confusing at a first glance, but it allows a very flexible management of raw and derived data in data acquisition. In order to understand it better, let's consider this common use case in data acquisition, where an array of raw samples is read and stored in the tree during data acquisition, together with the associated time values. In addition, converted data can be derived, e.g. by subtracting an offset value and mutiplying by a gain factor. If we want to keep in the pulse file both raw and converted data we may think that two different signals need to be stored, thus doubling the required space on disk. This is however not true, since, once raw data are available, only the knowledge of the gain and offset (two scalar values) is required to derive converted data. MDSplus expressions turn out to represent the ideal mean for expressing such dependency in data. We can convince ourselves making the following exercise miming a real data acquisition process (except for the actual dimension of the arrays):<br /> | + | This may seems a bit confusing at a first glance, but it allows a very flexible management of raw and derived data in data acquisition. In order to understand it better, let's consider this common use case in data acquisition, where an array of raw samples is read and stored in the tree during data acquisition, together with the associated time values. In addition, converted data can be derived, e.g. by subtracting an offset value and multiplying by a gain factor. If we want to keep in the pulse file both raw and converted data we may think that two different signals need to be stored, thus doubling the required space on disk. This is however not true, since, once raw data are available, only the knowledge of the gain and offset (two scalar values) is required to derive converted data. MDSplus expressions turn out to represent the ideal mean for expressing such dependency in data. We can convince ourselves making the following exercise miming a real data acquisition process (except for the actual dimension of the arrays):<br /> |
| In the first step we prepare an experiment model defining the following nodes: | In the first step we prepare an experiment model defining the following nodes: | ||
| * '''RAW_SAMPLES''': it will contain the raw samples | * '''RAW_SAMPLES''': it will contain the raw samples | ||
| Line 281: | Line 290: | ||
| TCL> put RAW_SAMPLES "[1,3,5,4,6,2,8,4,9,0]" | TCL> put RAW_SAMPLES "[1,3,5,4,6,2,8,4,9,0]" | ||
| Write times (10Hz starting at time 0) | Write times (10Hz starting at time 0) | ||
| - | TCL> put TIMES "[0.,0.1,0.2,0.3,0.4,0.5.0.6,0.7,0.8,0.9]" | + | TCL> put TIMES "[0.,0.1,0.2,0.3,0.4,0,5.0.6,0.7,0.8,0.9]" |
| At this point the pulse file contains the complete definition of both converted and raw signals, even if only the raw data array is actually stored in the pulse file. You may, for example, look at this signal with jScope (see next section). <br /> | At this point the pulse file contains the complete definition of both converted and raw signals, even if only the raw data array is actually stored in the pulse file. You may, for example, look at this signal with jScope (see next section). <br /> | ||
| The evaluation of expression SIGNAL2 yield in this case the array of converted signal values; evaluation of expression RAW_OF(SIGNAL2) yelds the array of raw samples and evaluation of expression DIM_OF(SIGNAL2) yields the array if sample times. | The evaluation of expression SIGNAL2 yield in this case the array of converted signal values; evaluation of expression RAW_OF(SIGNAL2) yelds the array of raw samples and evaluation of expression DIM_OF(SIGNAL2) yields the array if sample times. | ||
| Line 312: | Line 321: | ||
| Remember that TDI variable always begin with an underscore! </font> | Remember that TDI variable always begin with an underscore! </font> | ||
| - | <font face="Arial">You may wonder what are variables useful for. In fact, it makes no sense to use variable in insulated expressions, but expressions can be concatenated to form a TDI function, much like you would do in IDL, Matlab, or a programming language. Here is an example of a TDI function which returns the sum of all the members in a given array:</font> | + | <font face="Arial">You may wonder what are variables useful for. In fact, it makes no sense to use variable in insulated expressions, but expressions can be concatenated to form a TDI function, much like you would do in IDL, MATLAB, or a programming language. Here is an example of a TDI function which returns the sum of all the members in a given array:</font> |
| <font face="HE_TERMINAL">fun public vector_sum(in _a)<br /> {<br /> _sum = 0;<br /> for(_i = 0; _i < size(_a); _i++)<br /> _sum = _sum + _a[_i];<br /> return(_sum);<br /> }</font> | <font face="HE_TERMINAL">fun public vector_sum(in _a)<br /> {<br /> _sum = 0;<br /> for(_i = 0; _i < size(_a); _i++)<br /> _sum = _sum + _a[_i];<br /> return(_sum);<br /> }</font> | ||
| Line 351: | Line 360: | ||
| [1,2,3,4,5,6,7,8,9,10] | [1,2,3,4,5,6,7,8,9,10] | ||
| - | Things are now a bit more tricky...when evaluating expression SIGNAL2 (after opening the pulse file we used to put raw data and times in the previous examples) we may expect to get the signal samples, but we got a more complex thing. In fact the signal type is one of the types supported by MDSplus and is returned by evaluating expression SIGNAL2, that is, by reading the content of node SIGNAL2. It is however possible to force a further degree of evaluation using keyword '''data()'''. In this case the result will be a number either scalar or array. In case of SIGNAL2 it will be represented by the converted samples of the signal. The above example shows also how retrieving the actual raw samples and the associated times. | + | Things are now a bit more tricky...when evaluating expression SIGNAL2 (after opening the pulse file we used to put raw data and times in the previous example) we may expect to get the signal samples, but we got a more complex thing. In fact the signal type is one of the types supported by MDSplus and is returned by evaluating expression SIGNAL2, that is, by reading the content of node SIGNAL2. It is however possible to force a further degree of evaluation using keyword '''data()'''. In this case the result will be either a scalar or array. data(SIGNAL2) will yield the converted samples of the signal. Similarly the actual raw samples and the associated times arrays can be evaluated. |
| </font> | </font> | ||
| + | |||
| + | ---- | ||
| + | ===<center><font face="Arial"><big>Python integration in MDSplus expressions</big></font></center>=== | ||
| + | |||
| + | We shall see in the next sections that MDSplus is deeply integrated in python and all the MDSplus functionality is available when programming in python. The converse is also true, that is, it is possible to define python functions and use them in MDSplus expression.<br /> | ||
| + | We have seen in the previous section how a new function written in TDI can be defined so that it can be included in MDSplus expressions. We shall now do the same with a sample function '''my_python_sum''' written in python, making the sum of two numbers (scalar or arrays). | ||
| + | |||
| + | def my_python_sum(a,b): | ||
| + | return a+b | ||
| + | |||
| + | The python code must be saved in a file with the same name of the function (my_python_sum.py in this case) and in the directory referred by env variable MDS_PATH. This is the only required action to let MDSplus handle the python function in expressions. <br /> | ||
| + | We can exercise this python function in expression evaluation using tdic: | ||
| + | TDI> treeopen("my_tree",-1) | ||
| + | 265388041 | ||
| + | TDI> NUM2 | ||
| + | [1,2,3,4,5,6,7] | ||
| + | TDI> my_python_sum(2,3) | ||
| + | 5 | ||
| + | TDI> my_python_sum(2,[1,2,3]) | ||
| + | [3,4,5] | ||
| + | TDI> my_python_sum(2,NUM2) | ||
| + | [3,4,5,6,7,8,9] | ||
| + | |||
| + | We can do even more, that is, we can store in a MDSplus tree an expression which includes python function calls. Let's do it in my_tree: | ||
| + | TCL> set tree my_tree | ||
| + | TCL> put NUM3 "my_python_sum(NUM1, NUM2)" | ||
| + | |||
| + | And then let's evaluate NUM3 in tdic: | ||
| + | TDI> treeopen("my_tree",-1) | ||
| + | 265388041 | ||
| + | TDI> NUM1 | ||
| + | 2 | ||
| + | TDI> NUM2 | ||
| + | [1,2,3,4,5,6,7] | ||
| + | TDI> NUM3 | ||
| + | [3,4,5,6,7,8,9] | ||
| + | |||
| + | Recall that NUM3 does not contain the array itself, but the specification of a MDSplus expression implying the activation of the python function my_python_sum(). When evaluated, thw function is executed on the fly yielding the expected result. | ||
| ---- | ---- | ||
| Line 361: | Line 408: | ||
| ===<center><font face="Arial"><big>What next?</big></font></center>=== | ===<center><font face="Arial"><big>What next?</big></font></center>=== | ||
| - | <font face="Arial">We have now learnt how to build and populate a MDSplus tree. We know also how references into the tree are specified and we know that it is possible to use expressions for a flexible definition of data. </font> | + | <font face="Arial">We have now learnt how to build and populate a MDSplus tree and some basic concepts about MDSplus data. We know also how references into the tree are specified and we know that it is possible to use expressions for a flexible definition of data. </font> |
| <font face="Arial">We are now ready to go [[Documentation:Tutorial:DataAccess| further]], and to access data contained in the tree. The next tutorial will show you how to export MDSplus trees over the network, and how to access data from programs and from jScope, the MDSplus tool for waveform display.</font> | <font face="Arial">We are now ready to go [[Documentation:Tutorial:DataAccess| further]], and to access data contained in the tree. The next tutorial will show you how to export MDSplus trees over the network, and how to access data from programs and from jScope, the MDSplus tool for waveform display.</font> | ||
Current revision
|
[edit] General Notation: The Data Hierarchy - Trees, Nodes, and ModelsThe basic data structure of MDSplus is a self-descriptive hierarchy called a TREE. The hierarchy consists of large numbers of named NODES which make up the branches (structure) and leaves (data) of each tree. MDSplus SHOTS are trees created from a special type of tree called a MODEL, a template which contains all of the structure and setup data for an experiment or code. SHOTS are copies of the model augmented by the stored data and correspond to particular runs of an experiment or code. For a typical experiment, data from various sources are grouped in some logical manner and divided into a number of trees which each form the top level of their respective hierarchies. These trees themselves can be organized into a hierarchy with a root tree and SUBTREES as in the following figure.
[edit] How to create and populate a MDSplus treeIn this first lesson we shall build a very simple MDSplus tree. A MDSPlus tree is a database which contains several types of data. A data item may be a number, a string, a signal or, more generally an expression i.e. a combination of data (possibly stored in the same tree) and operators. The MDSplus tree represents the core of MDSplus, and there are many ways for interacting with it. Here, we shall use two tools: 1) The Tree Command Language (TCL, not to be confused with TCL/TK) for creating and editing a tree; 2) The jTraverser tool for providing a graphical interface to the tree. [edit] A Quick Check for some important environment variablesBefore starting working with MDSplus, it is important to make sure that a few environment variables are properly set. When installed via an install shield or RPM, the variables should be ok, but in the case MDSplus is build from the source distribution, the following variables must be defined for Linux:
[edit]
|